mainmenu
- About Center
- Members
- Research
- Publications
- Facilities
- Super-depth Optical Imaging Lab
- Ultrafast Infrared Spectroscopy Lab
- Ultrafast IR-Visible Spectroscopy Lab
- Instrument and Chemistry Lab
- Molecular Imaging Lab
- Multidimensional Infrared Spectroscopy Lab
- Sample Preparation Lab
- Optical Frequency Comb Spectroscopy Lab
- Single Molecule Imaging Lab
- Spectroscopic Imaging Lab
- Theory and Computation
- Open Access Facility
- Collaborative Research Equipment
- Quantum Spectroscopy Lab
- Ultrafast Raman Probe Spectroscopy Lab
- Equipment Inventory
- Board
Facilities
- Super-depth Optical Imaging Lab
- Ultrafast Infrared Spectroscopy Lab
- Ultrafast IR-Visible Spectroscopy Lab
- Instrument and Chemistry Lab
- Molecular Imaging Lab
- Multidimensional Infrared Spectroscopy Lab
- Sample Preparation Lab
- Optical Frequency Comb Spectroscopy Lab
- Single Molecule Imaging Lab
- Spectroscopic Imaging Lab
- Theory and Computation
- Open Access Facility
- Collaborative Research Equipment
- Quantum Spectroscopy Lab
- Ultrafast Raman Probe Spectroscopy Lab
Theory and Computation
Cluster-type computer
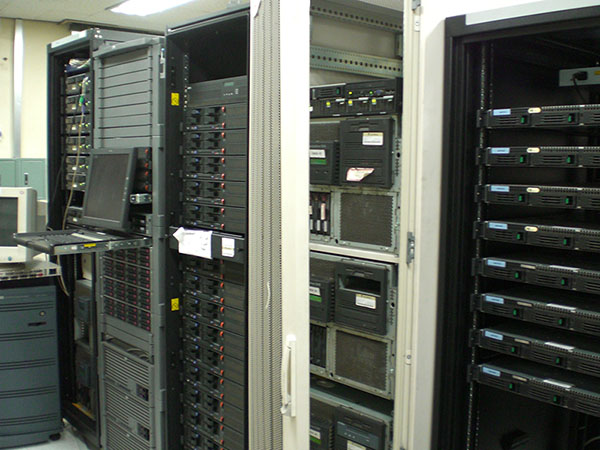
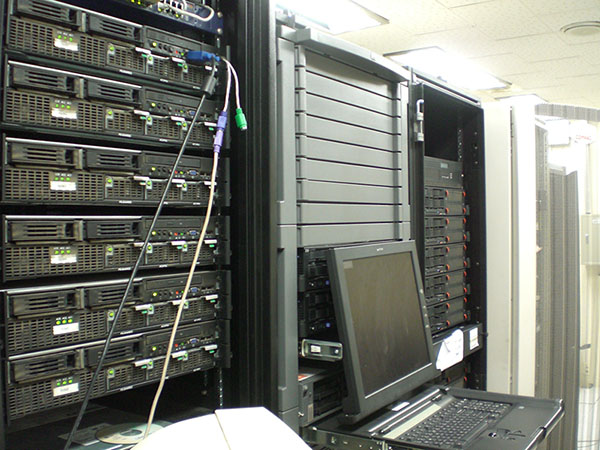


Work station
Cluster | Nodes | CPU/GPU | S:C;T/Ga | Memory per node | Slurm partition |
|---|---|---|---|---|---|
alpha[1-4] | 4 | Intel Xeon E5-2630 v3 (2.4 GHz Haswell) | 2:8:2 | 128 GB | alpha-1 |
alpha[5-8] | 4 | Intel Xeon E5-2630 v3 (2.4 GHz Haswell) | 2:8:2 | 128 GB | alpha-2 |
alphpha[9-44] | 36 | Intel Xeon E5-2680 v3 (2.5 GHz Haswell) | 3:12:2 | 128 GB | alpha-3 |
beta[1-48] | 48 | Intel Xeon E3-1241 v3 (3.5 GHz Haswell) | 1:4:2 | 32 GB | beta |
gamma[1-4] | 4 | Intel Xeon 5160 (3.0 GHz Woodcrest) | 2:2:1 | 4 GB | gamma-1 |
gamma[5-8] | 4 | Intel Xeon 5160 (3.0 GHz Woodcrest) | 2:2:1 | 8 GB | gamma-2 |
delta[1-16] | 16 | Intel xeon E5430 (2.66 GHz Harpertown) | 2:4:1 | 8 GB | delta |
eta[1-2] | 2 | Intel Core i7-2600 (3.4 GHz Sandy Bridge)/ Nvidia GTX 1080 | 1:4:2/1 | 16 GB | etacpu-1 etagpu-1 |
eta[3-26] | 24 | AMD Ryzen 1700 (3.0 GHz Zen)/ Nvidia GTX 1080 Ti | 1:8:2/1 | 32 GB | etacpu-2 etacpu-2 |
theta[1-4] | 4 | Intel Xeon Gold 6230 (2.1 GHz Cascade Lake) | 4:20:2 | 384 GB | theta |
eps[1-4] | 4 | Intel Xeon E5-2640 (2.5 GHz Sandy Bridge EP) | 2:6:2 | 64 GB | eps |
zeta[1-8] | 8 | Intel Xeon E3 1270 (3.5 GHz Ivy Bridge) | 1:4:2 | 16 GB | zeta |
Storage
Name | CPU | RAM | HDD | Raid Type | OS |
|---|---|---|---|---|---|
Damore1 | P4 2.4GHZ | 512MB | 120GB * 8 | 5 | >RedHat 8.0 |
Damore2 | P4 2.4GHZ | 512MB | 120GB * 8 | 5 | >RedHat 8.0 |
Damore3 | P4 2.4GHZ | 512MB | 120GB * 4 + 250GB * 4 | 5 | >RedHat 8.0 |
Damore4 | P4 2.4GHZ | 512MB | 250GB * 8 | 5 | >RedHat 8.0 |
Damore5 | P4 2.4GHZ | 512MB | 250GB * 8 | 5 | >RedHat 8.0 |
ADTX1 |
|
| 250GB * 8 | 5 |
|
ADTX2 |
|
| 250GB * 8 | 5 |
|
Software
company | Name | Ver. | Purpose |
|---|---|---|---|
Gaussian | Gaussian | 98 A.6, A.7, A.8, A.9 G03, C02, D01, G16 | Ab initio calculation |
Gaussian | Gauss view |
| Modeling and visualizing |
Gaussian | Linda | 3.1 | Parallelizing of Gaussian |
Fujitsu | WinMopac Mopac2002 | 2.0 | Molecular Dynamics calculation |
Gamess | GAMESS |
| Ab initio calculation |
SGI | Cerius 2 | Release 4.0 | Modeling and visualizing + Ab initio calculation |
Cambridge | ChemOffice Ultra | 6.0 | Modeling and visualizing |
Hypercube | HyperChem | 5.02 | Modeling and visualizing |
Amber | Amber | 7 | Molecular Dynamics calculation |
Charmm | Charmm | 30b2 | Hybrid calculation of Ab initio and MD |